Project Tag: featured
-
Fast4DReg: Fast registration of 4D microscopy datasets

Unwanted sample drift is a common issue that plagues microscopy experiments, preventing accurate temporal visualization and quantification of biological processes. Although multiple methods and tools exist to correct images post-acquisition, performing drift correction of three-dimensional (3D) videos using open-source solutions remains challenging and time-consuming. Here, we present a new tool developed for ImageJ or Fiji…
-
MYO10-filopodia support basement membranes at pre-invasive tumor boundaries
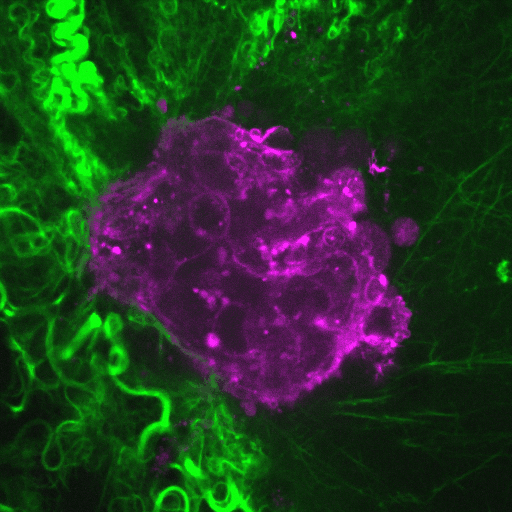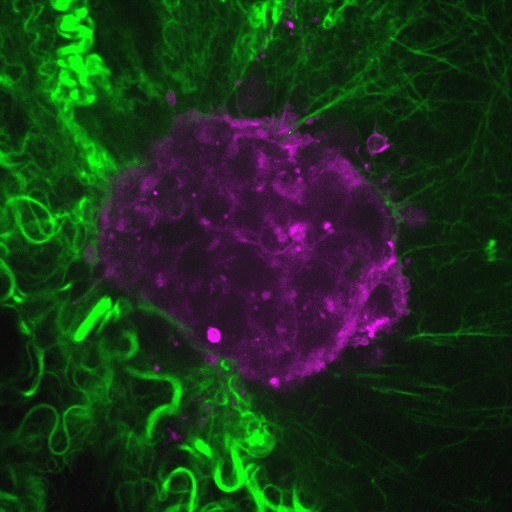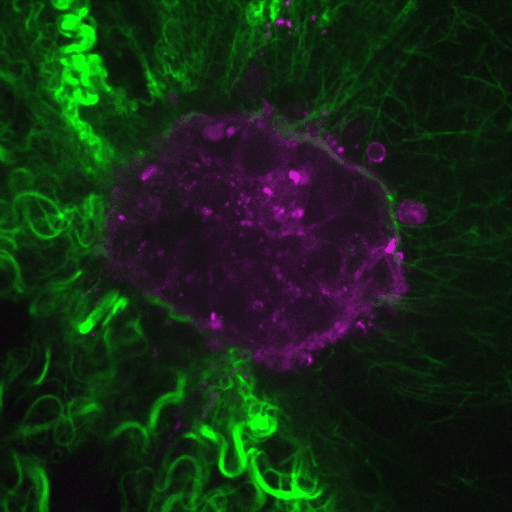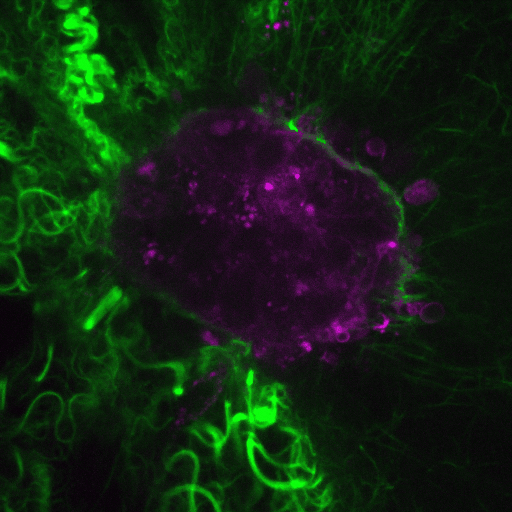
Ductal carcinoma in situ (DCIS) is a pre-invasive stage of breast cancer. During invasion, the encapsulating DCIS basement membrane (BM) is compromised, and tumor cells invade the surrounding stroma. The mechanisms that regulate functional epithelial BMs in vivo are poorly understood. Myosin-X (MYO10) is a filopodia-inducing protein associated with metastasis and poor clinical outcome in…
-
Myosin-X recruits lamellipodin to filopodia tips
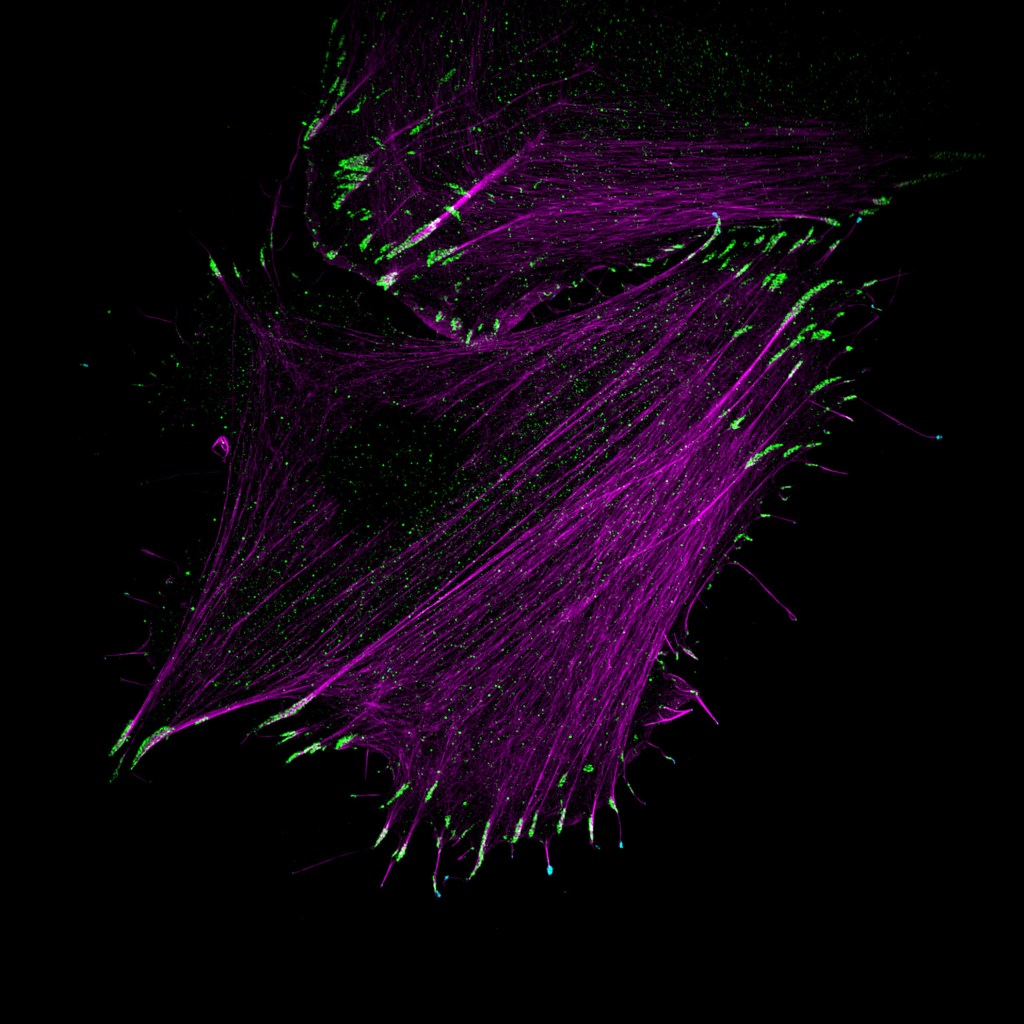
Myosin-X (MYO10), a molecular motor localizing to filopodia, is thought to transport various cargo to filopodia tips, modulating filopodia function. Yet, only a few MYO10 cargoes have been described. Here, using GFP-Trap and BioID approaches combined with mass spectrometry, we identified lamellipodin (RAPH1) as a novel MYO10 cargo. We report that the FERM domain of…
-
TrackMate 7: integrating state-of-the-art segmentation algorithms into tracking pipelines

TrackMate is an automated tracking software used to analyze bioimages and is distributed as a Fiji plugin. Here, we introduce a new version of TrackMate. TrackMate 7 is built to address the broad spectrum of modern challenges researchers face by integrating state-of-the-art segmentation algorithms into tracking pipelines. We illustrate qualitatively and quantitatively that these new…
-
TLNRD1 is a novel actin-bundling protein that promotes filopodia formation
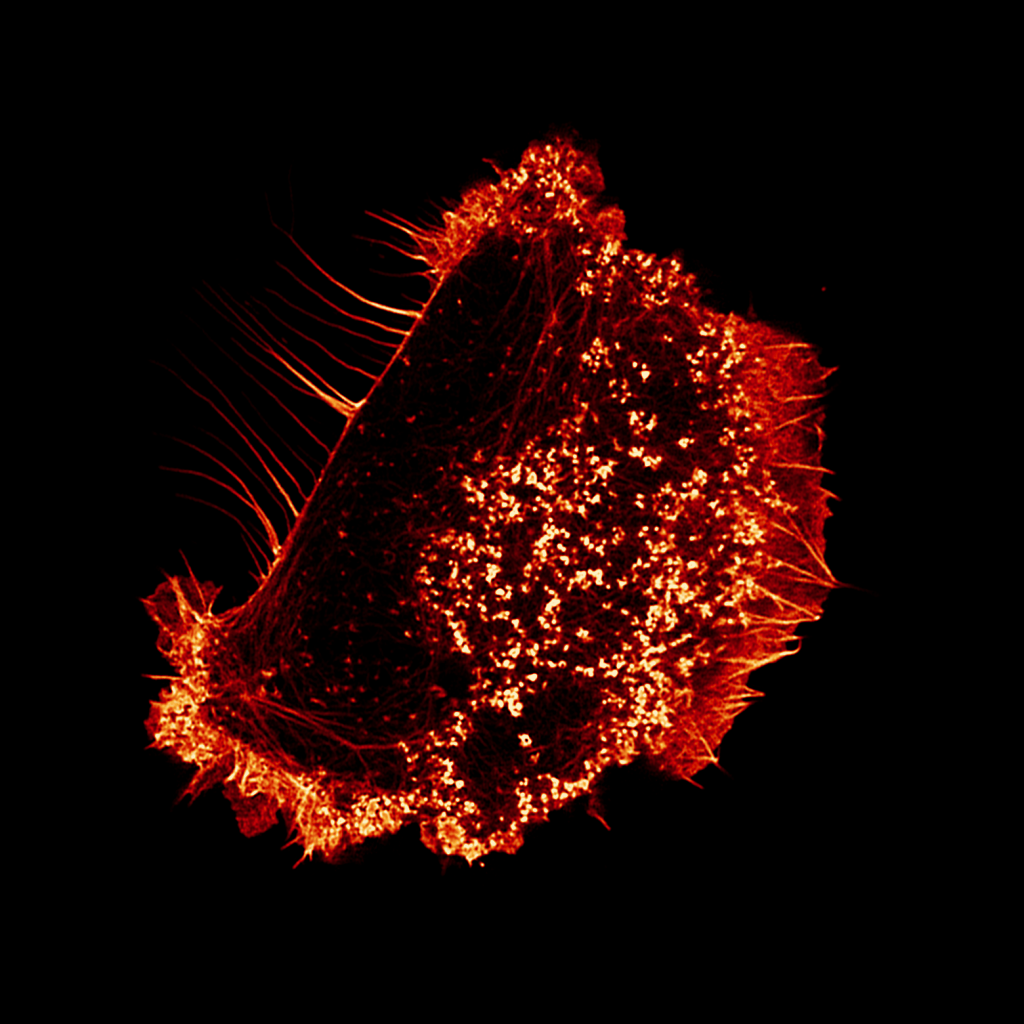
Talin is a mechanosensitive adapter protein that couples integrins to the cytoskeleton. Talin rod domain–containing protein 1 (TLNRD1) shares 22% homology with the talin R7R8 rod domains and is highly conserved throughout vertebrate evolution, although little is known about its function. We show that TLNRD1 is an α-helical protein structurally homologous to talin R7R8. Like…
-
Myosin-X and talin modulate integrin activity at filopodia tips
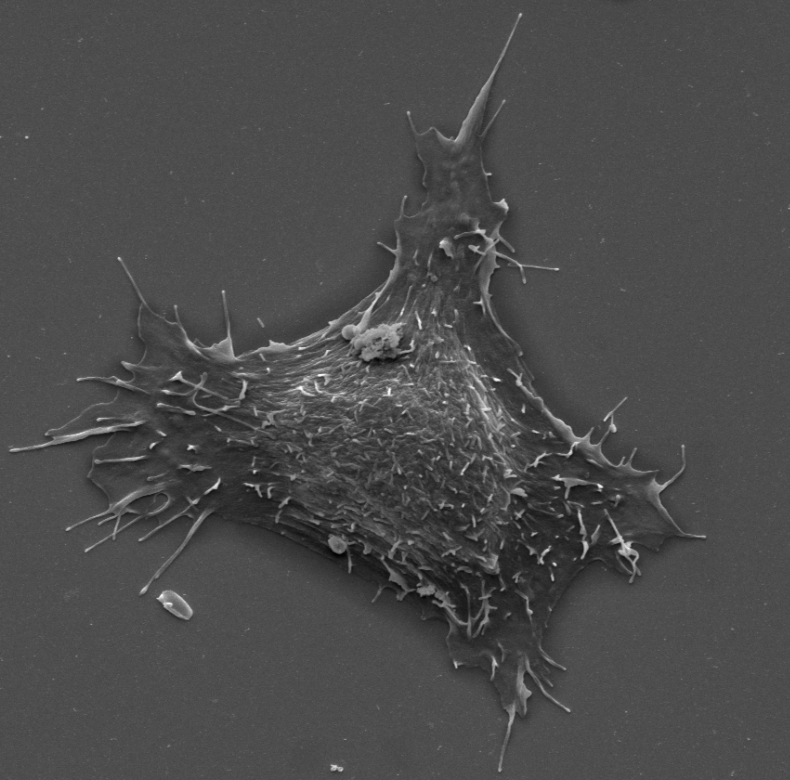
Filopodia assemble unique integrin-adhesion complexes to sense the extracellular matrix. However, the mechanisms of integrin regulation in filopodia are poorly defined. Here, we report that active integrins accumulate at the tip of myosin-X (MYO10)-positive filopodia, while inactive integrins are uniformly distributed. We identify talin and MYO10 as the principal integrin activators in filopodia. In addition,…
-
Democratising deep learning for microscopy with ZeroCostDL4Mic

Deep Learning (DL) methods are powerful analytical tools for microscopy and can outperform conventional image processing pipelines. Despite the enthusiasm and innovations fuelled by DL technology, the need to access powerful and compatible resources to train DL networks leads to an accessibility barrier that novice users often find difficult to overcome. Here, we present ZeroCostDL4Mic,…
-
Automated cell tracking using StarDist and TrackMate

The ability of cells to migrate is a fundamental physiological process involved in embryonic development, tissue homeostasis, immune surveillance, and wound healing. Therefore, the mechanisms governing cellular locomotion have been under intense scrutiny over the last 50 years. One of the main tools of this scrutiny is live-cell quantitative imaging, where researchers image cells over…
-
Cell matrix adhesion in cell migration

The ability of cells to migrate is a fundamental physiological process involved in embryonic development, tissue homeostasis, immune surveillance and wound healing. In order for cells to migrate, they must interact with their environment using adhesion receptors, such as integrins, and form specialized adhesion complexes that mediate responses to different extracellular cues. In this review,…
-
Filopodia in cell adhesion, 3D migration and cancer cell invasion
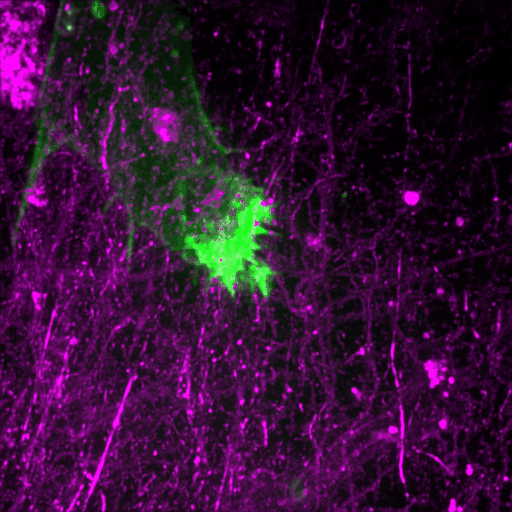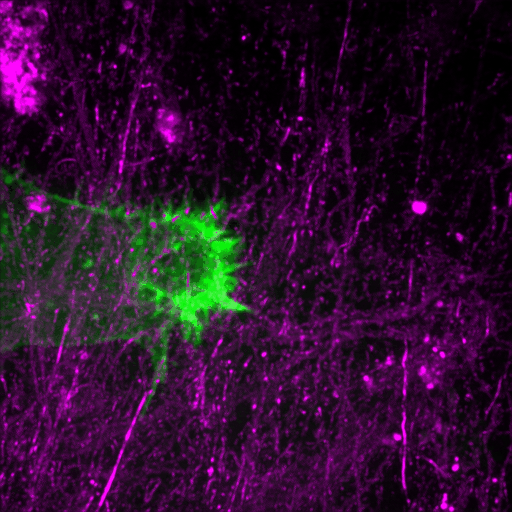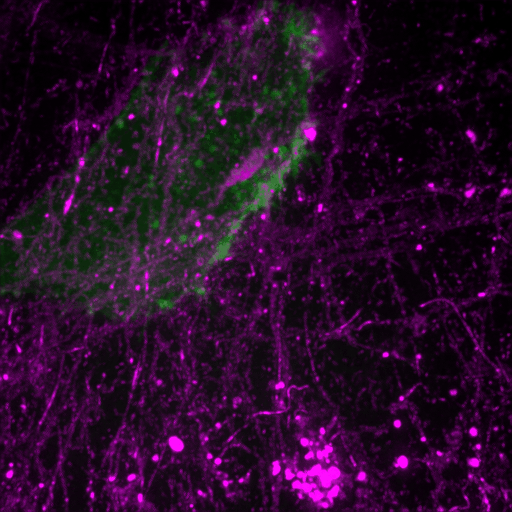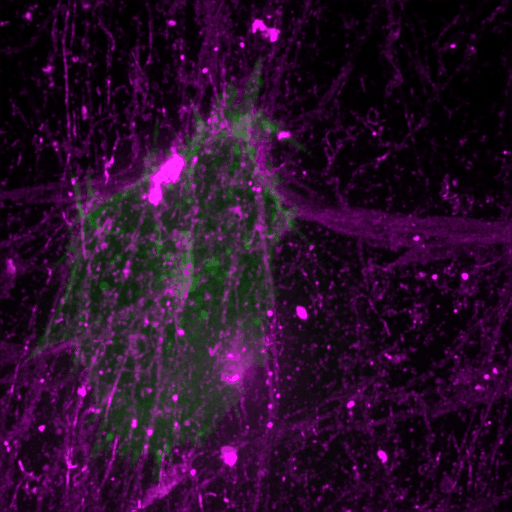
This review discusses recent advances in our understanding of the role filopodia and filopodia-like structures in cell adhesion and three dimensional (3D) cell migration both in vitro and in vivo. In particular, we focus on recent advances demonstrating that filopodia are involved in substrate tethering and environment sensing in vivo. We further discuss the emerging role of filopodia and…
-
Fluctuation-Based Super Resolution Traction Force Microscopy

Cellular mechanics play a crucial role in tissue homeostasis and are often misregulated in disease. Traction force microscopy is one of the key methods that has enabled researchers to study fundamental aspects of mechanobiology; however, traction force microscopy is limited by poor resolution. Here, we propose a simplified protocol and imaging strategy that enhances the…
-
Produce cell-derived matrices for migration studies
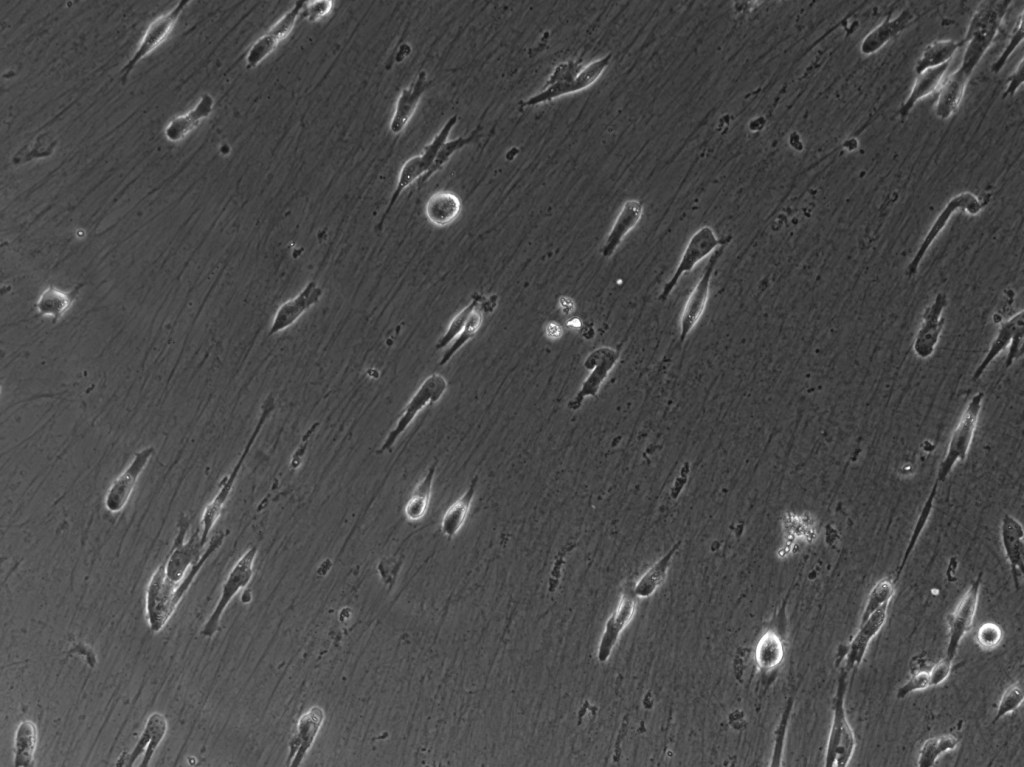
Cell-derived matrices for studying cell proliferation and directional migration in a complex 3D microenvironment 2D surfaces offer simple analysis of cells in culture, yet these often yield different cell morphologies and responses from those observed in vivo. Considerable effort has therefore been expended on the generation of more tissue-like environments for the study of cell…
-
Tracking filopodia using FiloQuant
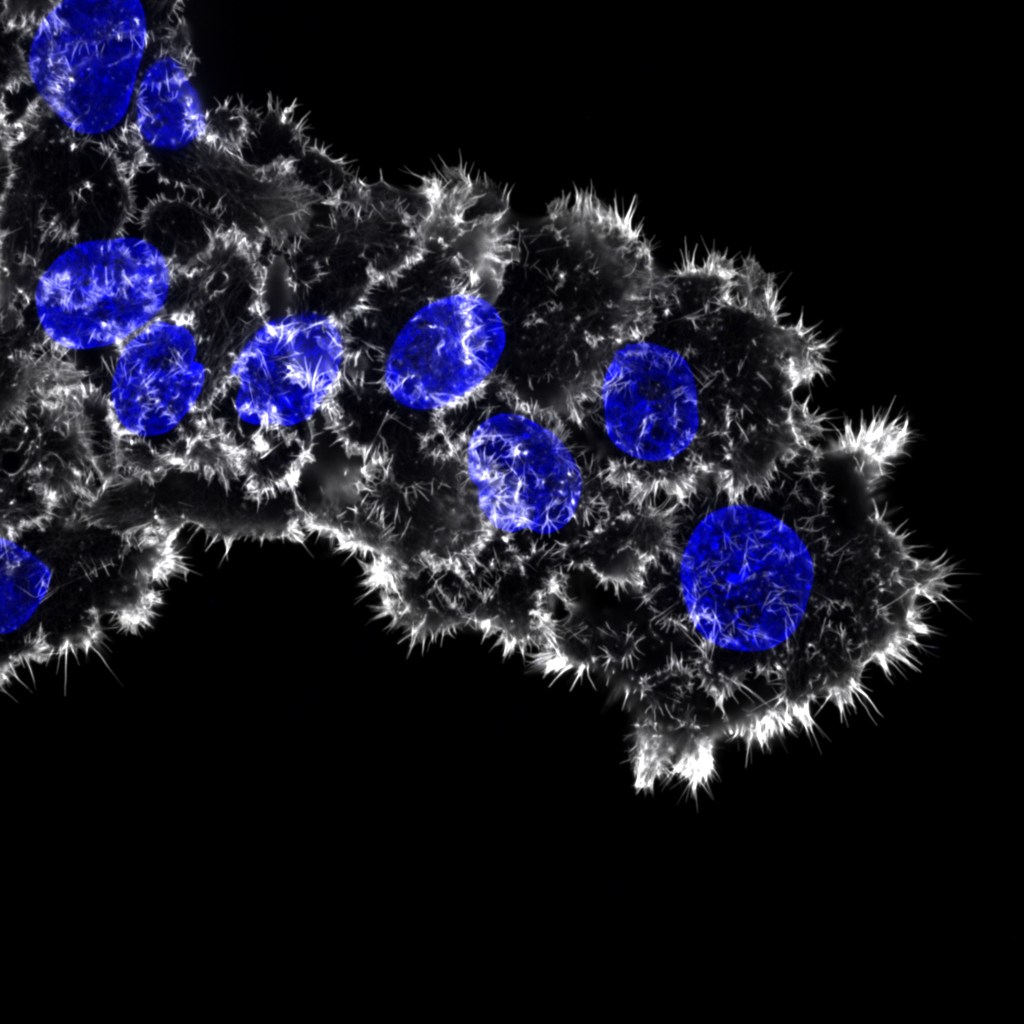
FiloQuant reveals increased filopodia density during breast cancer progression Defective filopodia formation is linked to pathologies such as cancer, wherein actively protruding filopodia, at the invasive front, accompany cancer cell dissemination. Despite wide biological significance, delineating filopodia function in complex systems remains challenging and is particularly hindered by lack of compatible methods to quantify filopodia properties. Here,…
-
Targeting filopodia to block cancer cell invasion

L-type calcium channels regulate filopodia stability and cancer cell invasion downstream of integrin signalling Mounting in vitro, in vivo and clinical evidence suggest an important role for filopodia in driving cancer cell invasion. Using a high-throughput microscopic-based drug screen, we identify FDA-approved calcium channel blockers (CCBs) as potent inhibitors of filopodia formation in cancer cells.…
-
Mapping the Filopodome
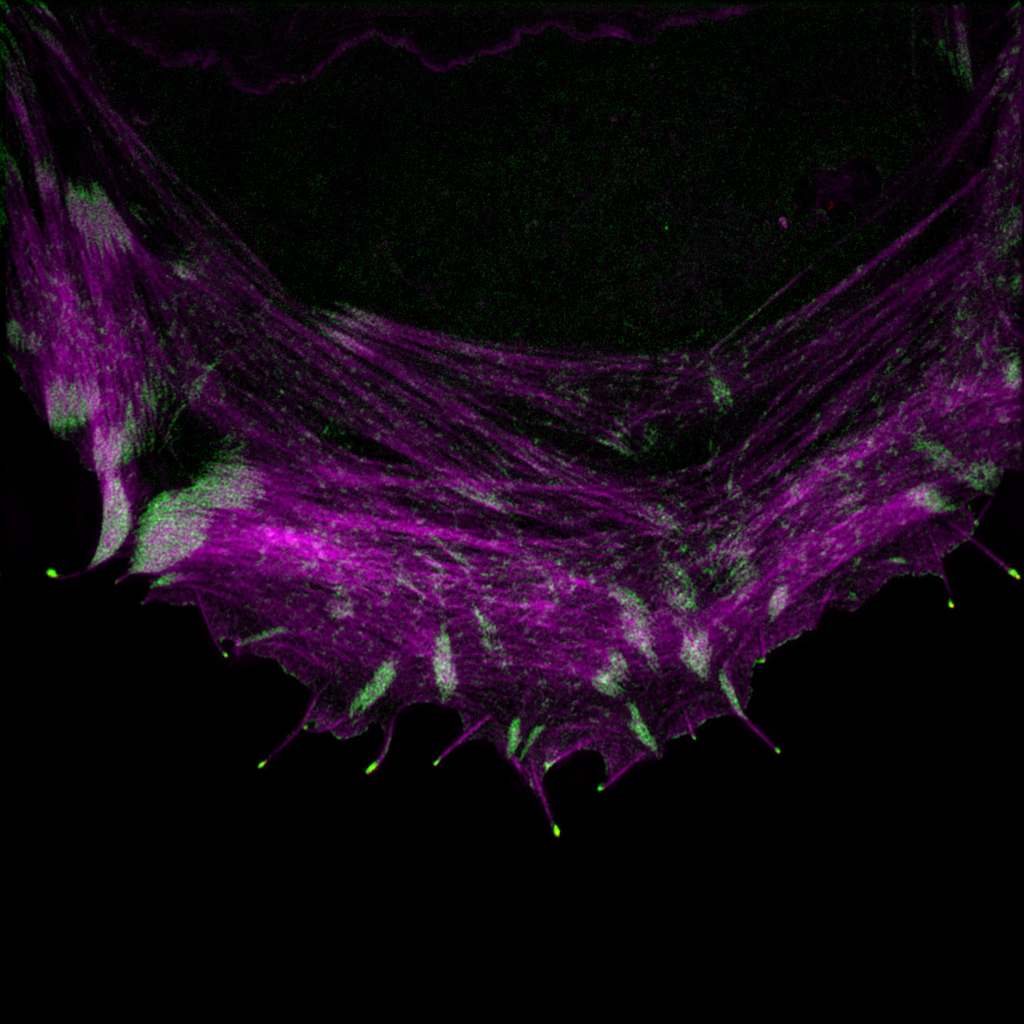
Filopodome Mapping Identifies p130Cas as a Mechanosensitive Regulator of Filopodia Stability Filopodia are adhesive cellular protrusions specialized in the detection of extracellular matrix (ECM)-derived cues. Although ECM engagement at focal adhesions is known to trigger the recruitment of hundreds of proteins (“adhesome”) to fine-tune cellular behavior, the components of the filopodia adhesions remain undefined. Here,…