Our Research interests
Cancer metastasis
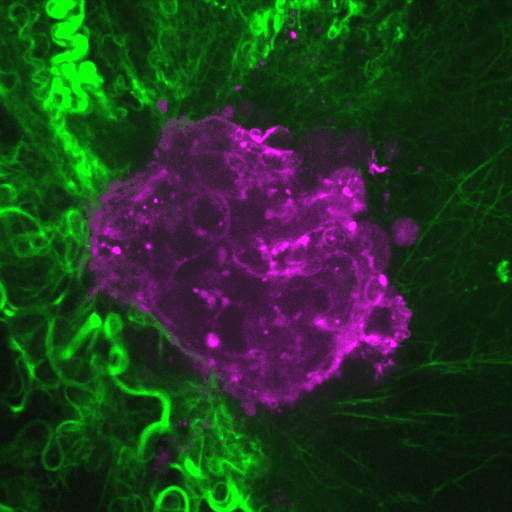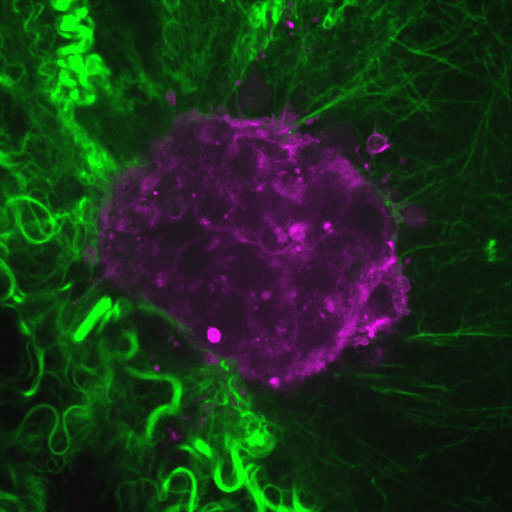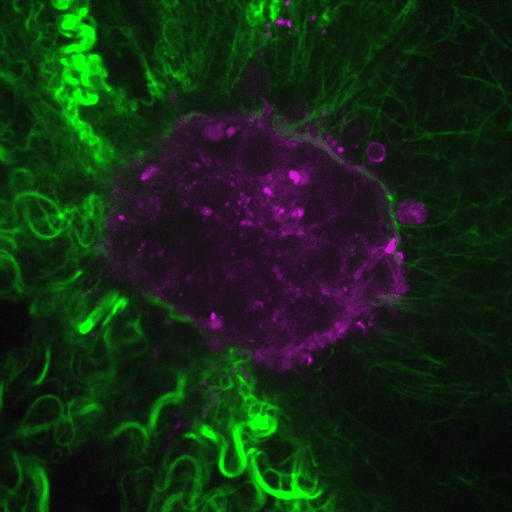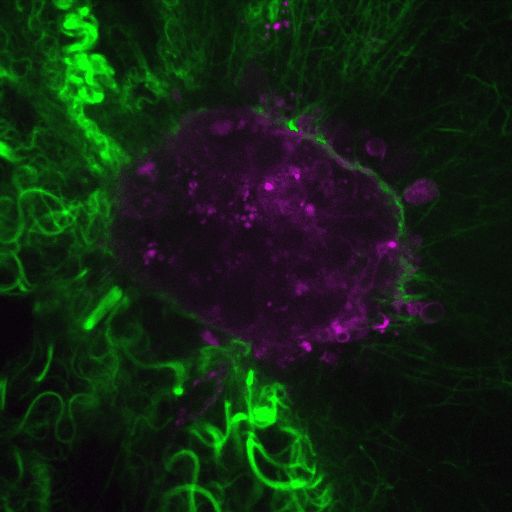
Our research aims to elucidate the mechanisms by which cancer cells spread through the body.
Cancer is a leading cause of death worldwide and accounts for 8.2 million deaths per year.
The formation of metastases is responsible for 90% of deaths in patients with solid tumors.
Video: Cancer cells (purple) invade through collagen (green). Credit : Guillaume Jacquemet and Emilia Peuhu
Cell migration
Our research aims to elucidate the molecular mechanisms by which cells move.
Cell migration is a fundamental physiological process involved in embryonic development, tissue homeostasis, immune surveillance, and wound healing.
For cells to migrate, they must interact with their environment using adhesion receptors, such as integrins, and form specialized adhesion complexes that mediate responses to different extracellular cues.
Video: Cancer cell (green) migrating on cell-derived matrices (magenta). Credit : Guillaume Jacquemet
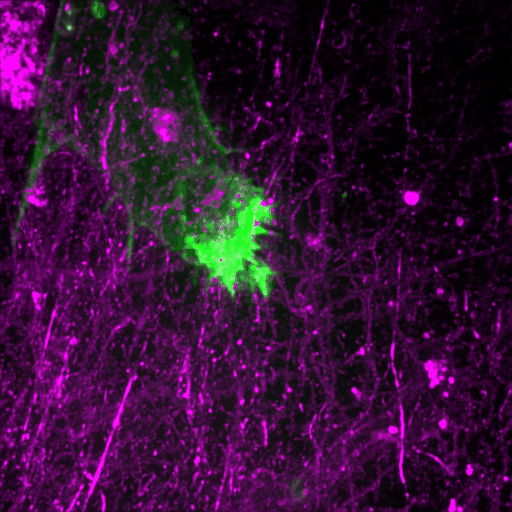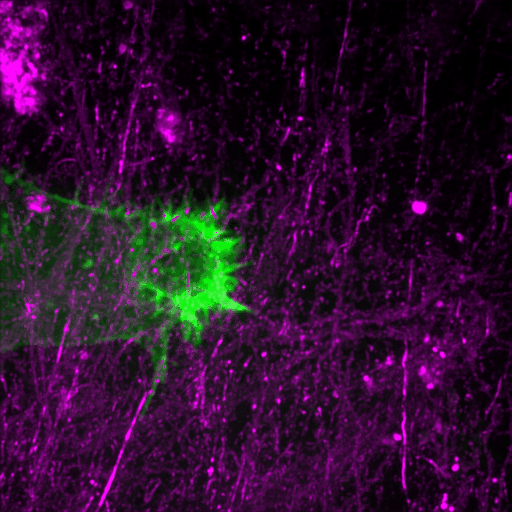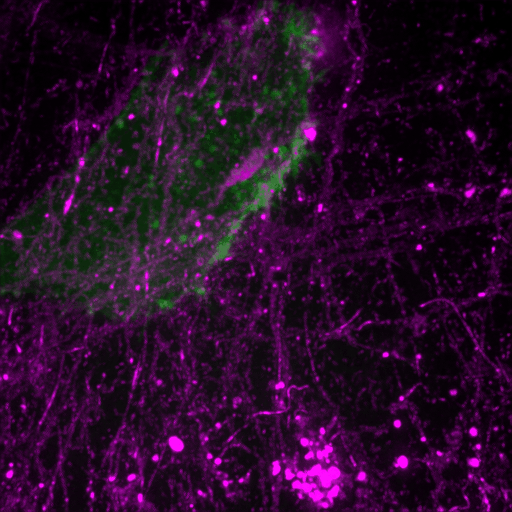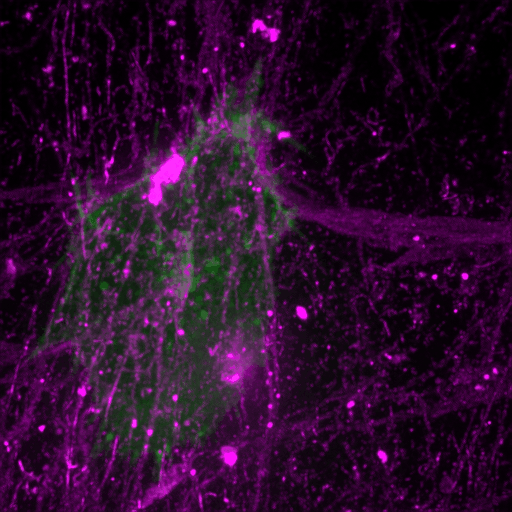
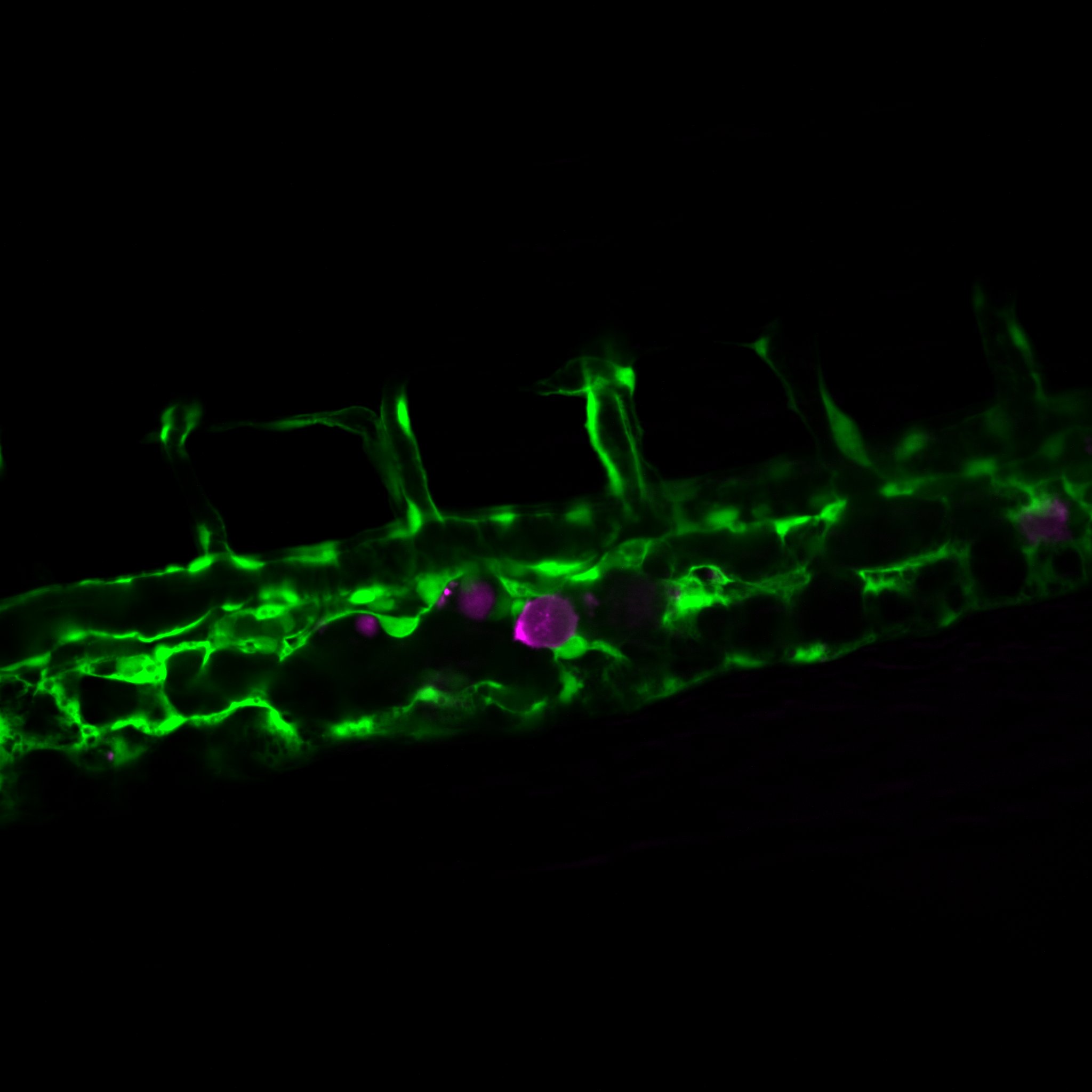
How cancer cells and immune cells interact with the vasculature
We study how cancer cells and immune cells interact with blood vessels because these encounters strongly influence how tumors grow, spread, and respond to treatment.
Using microfluidic “vessel-on-a-chip” systems, we can recreate tiny blood vessels in the lab and watch cells move and communicate in a controlled, lifelike environment. With transparent zebrafish embryos, we can observe these same processes inside a living body in real time.
As part of of the center of excellence IMMENs we also study lymphatics.
Picture: A cancer cell (magenta) inside the vasculature of a zebrafish embryo. Credit : Guillaume Jacquemet
Filopodia
Our research aims to unravel how cells can sense their environment.
Filopodia are actin-rich “antenna-like” protrusions that are responsible for constantly probing the cellular environment.
In cancer cells, filopodia contribute to single cell migration and invasion, and filopodia-like structures have been implicated in cancer cell survival at metastatic sites.
Picture: Cancer cell labeled for F-actin. Credit : Guillaume Jacquemet
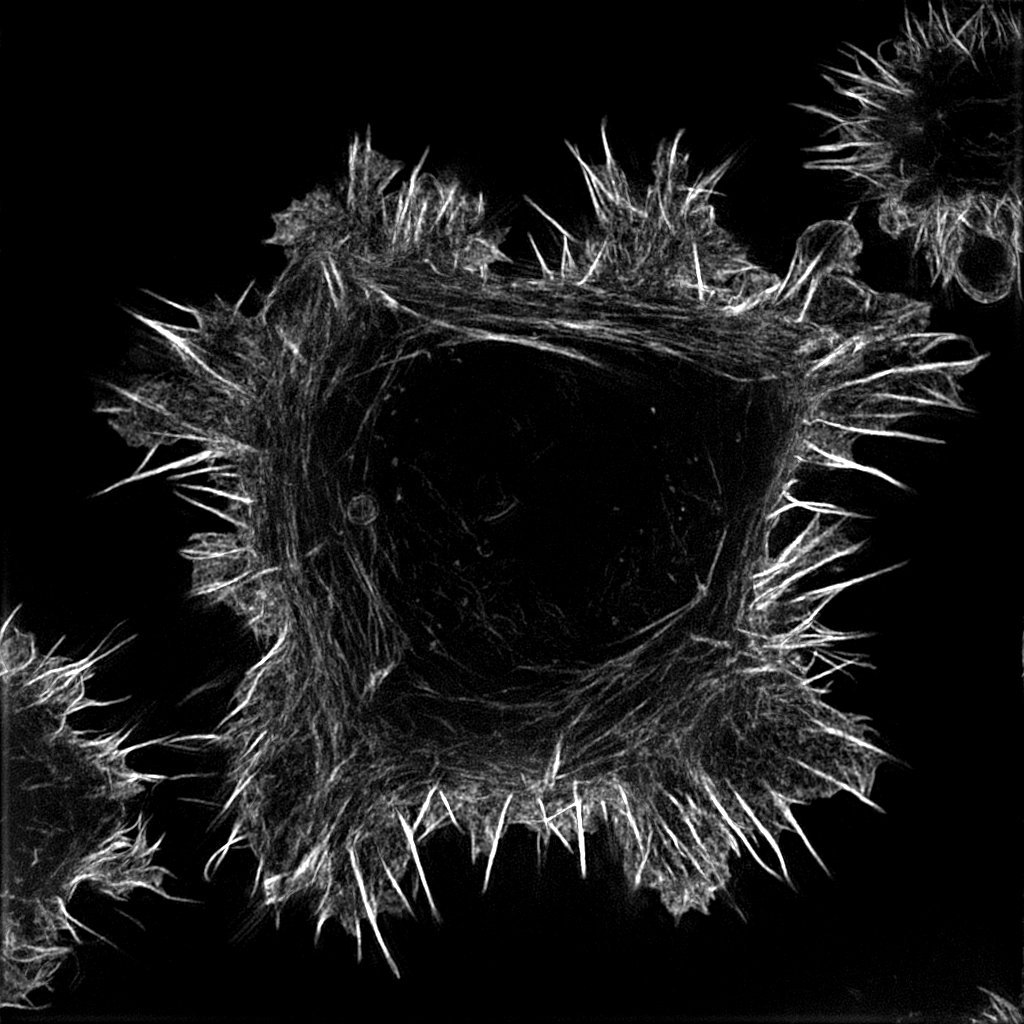
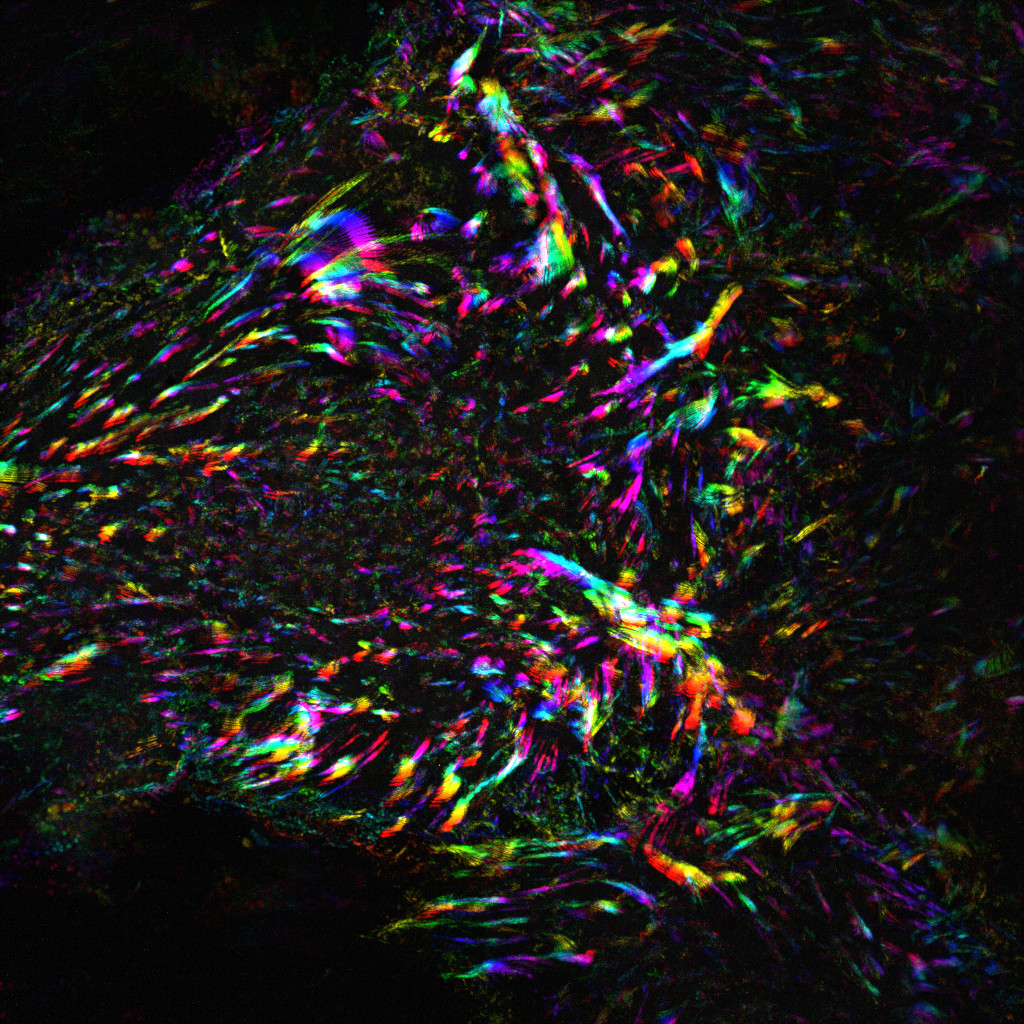
Image analysis
We use multiple types of microscopy approaches in our work and continuously develop tools to analyze images.
Check our software.
Image: Time projection illustrating the dynamics of focal adhesions in human endothelial cells. Credit : Martina Lerche and Guillaume Jacquemet
You must be logged in to post a comment.