Deep learning to analyse microscopy images
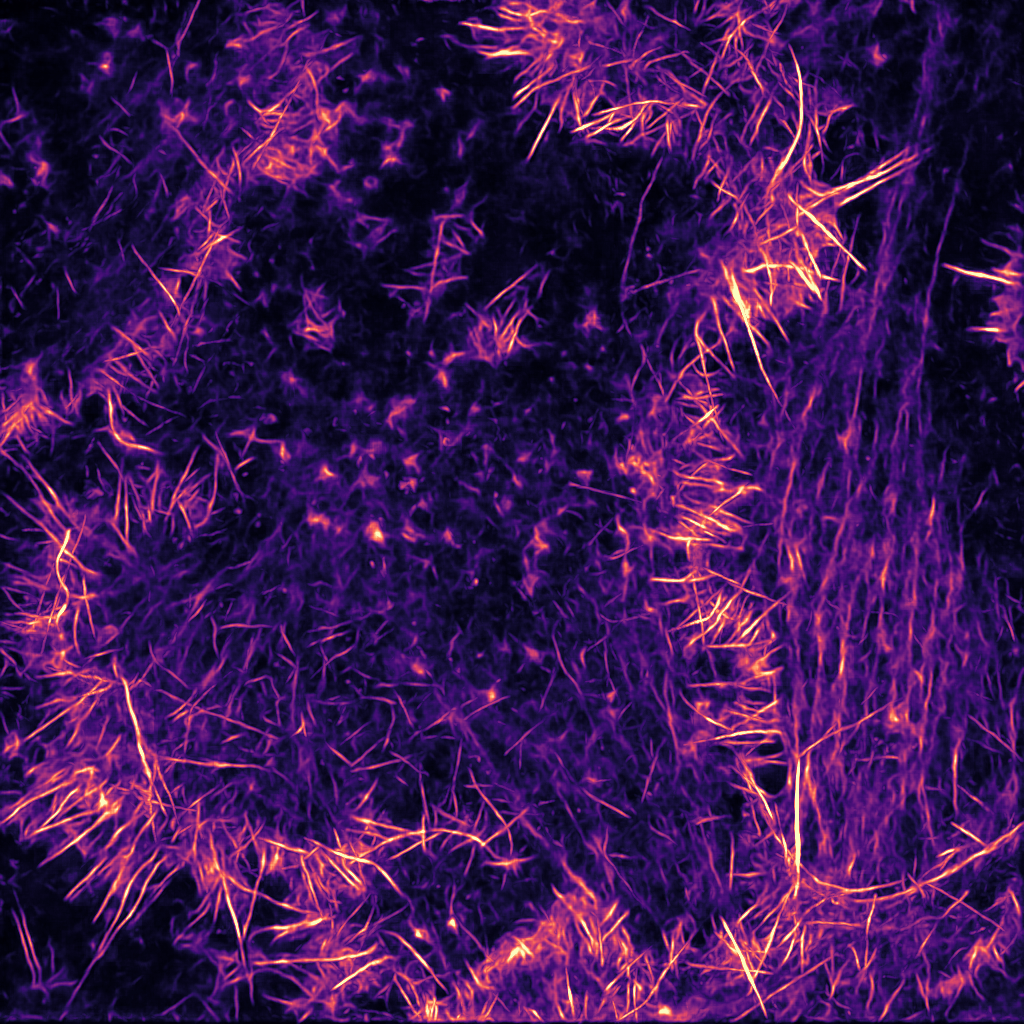
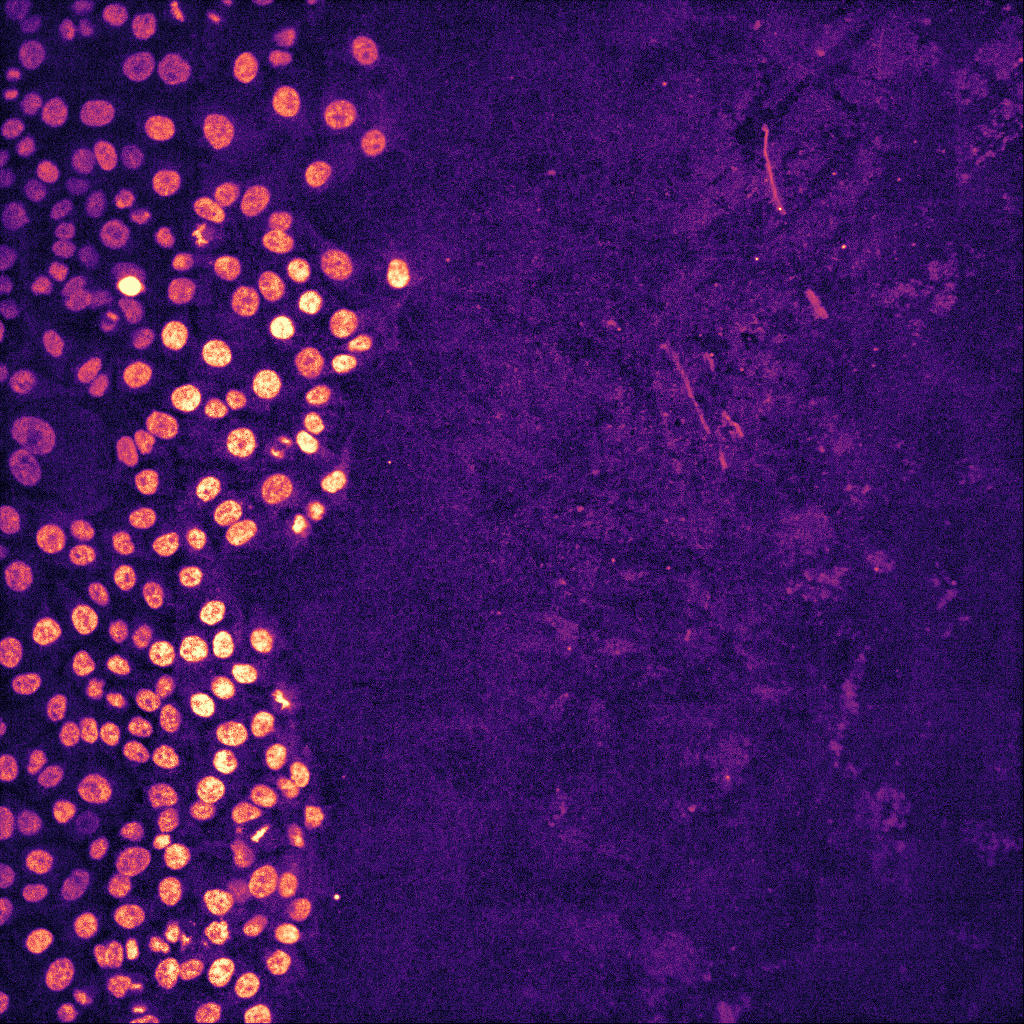
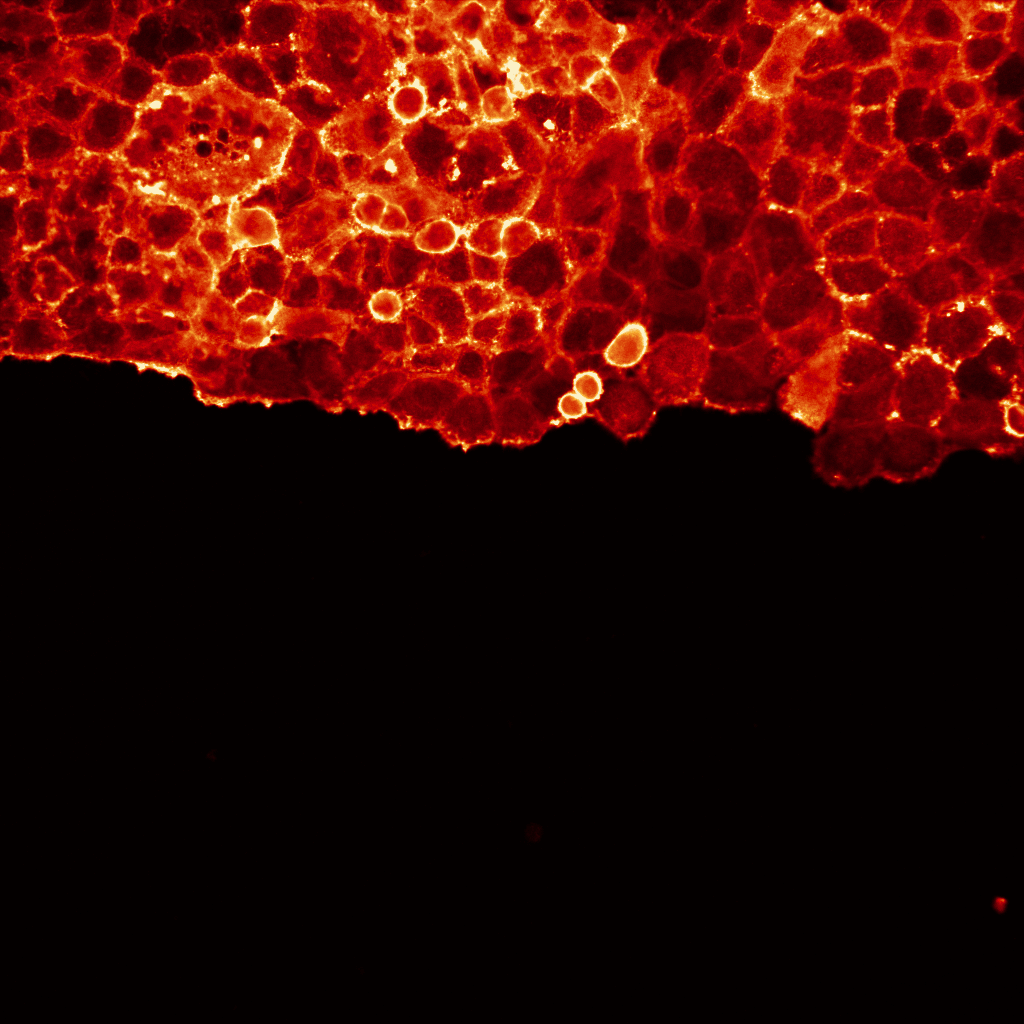
Artificial intelligence (AI)-powered algorithms are now influencing many aspects of our day-to-day life, from providing movies/music recommendations to controlling self-driving cars. These algorithms are also increasingly used in the lab to aid biomedical research. In particular, the ability to analyse and process images using AI is slowly revolutionizing the quality and quantity of data we collect from microscopy images. In fact, AI-based algorithms can now be applied to perform virtually any high-performance image analysis tasks such as classifying images, detecting and segmenting objects, aligning images or improving image quality by removing noise or increasing image resolution.
Read our primer on “Deep learning to analyse microscopy images” published in The Biochemist.
Democratising deep learning for microscopy with ZeroCostDL4Mic
ZeroCostDL4Mic is an entry-level platform simplifying DL access by leveraging the free, cloud-based computational resources of Google Colab. ZeroCostDL4Mic allows researchers with no coding expertise to train and apply key DL networks to perform tasks including segmentation (using U-Net and StarDist), object detection (using YOLOv2), denoising (using CARE and Noise2Void), super-resolution microscopy (using Deep-STORM), and image-to-image translation (using Label-free prediction – fnet, pix2pix, and CycleGAN). Importantly, we provide suitable quantitative tools for each network to evaluate model performance, allowing model optimization.
Read our ZeroCostDL4Mic paper here
Access the ZeroCostDL4Mic platform here.